For the letter T in our A-Z series, Chris discusses the emerging field of Therapygenetics.
In a nutshell (which therapy is most likely to work for whom and why?)
Imagine how great it would be if you knew how likely a treatment was to work for you before embarking on it. That is what this field is all about.
Just like many human traits (including anxiety, learning ability and response to the environment), response to psychological therapy is likely to be influenced by genetics. Therapygenetics research focuses on detecting clinically useful genetic variation associated with therapy response. This is so we can use genetic information in prediction models (alongside patient data), to predict which treatment type is most likely to work for whom i.e. personalised medicine. We also hope that by identifying genes associated with therapy response, we can understand more about the underlying biology. This work is in its early stages. However, progress in psychiatry, genetics and improved access to psychological therapy will provide us with the methods and sample sizes needed to make a clinical breakthrough.
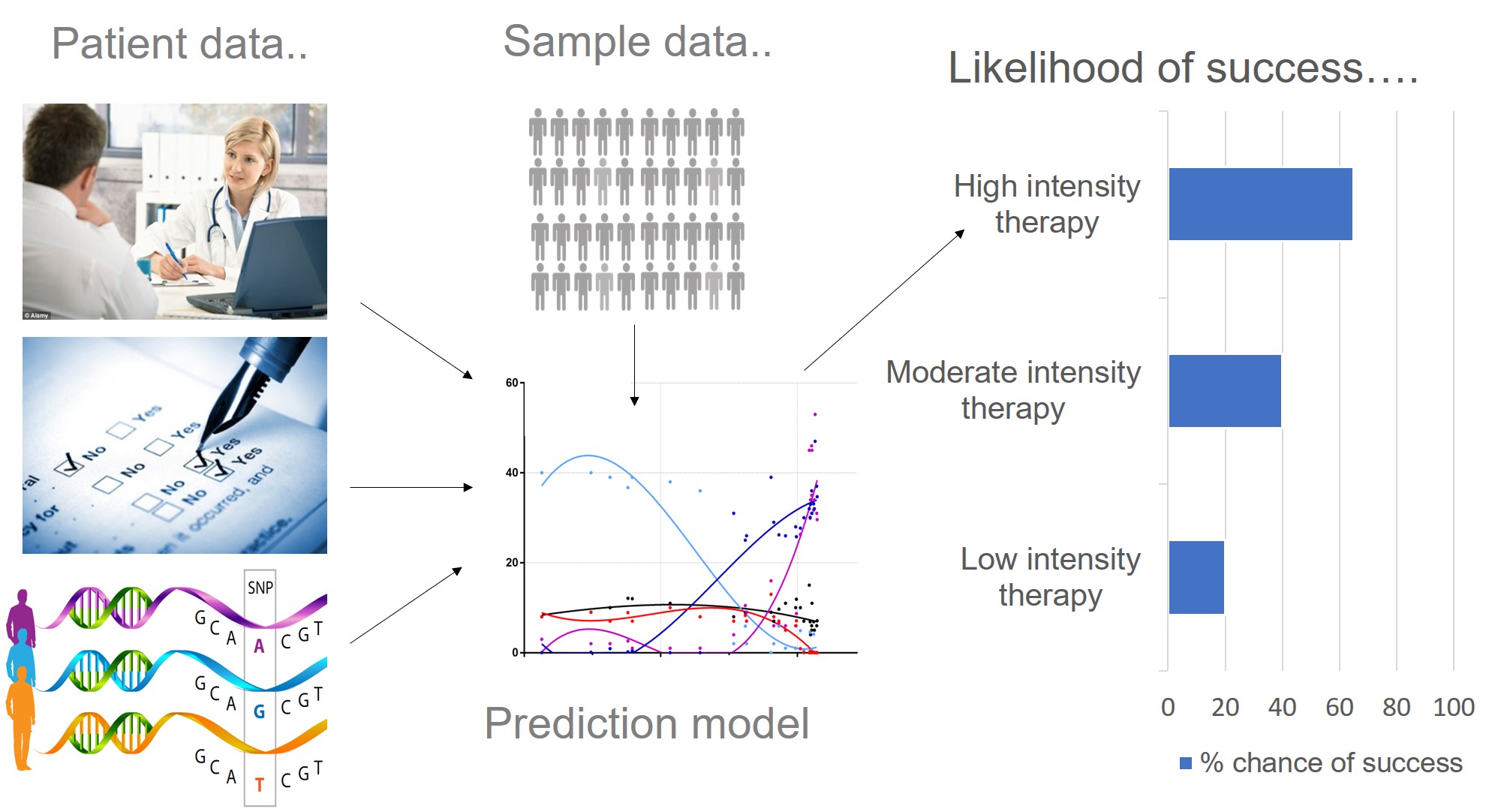
Why do we think predicting treatment response is important?
Anxiety and depressive symptoms and disorders are common. They affect at least 1 in 4 people in their lifetime and are often chronic and disabling (Steel et al. 2014). These disorders are a leading cause of disability worldwide and cost the NHS tens of billions each year (and rising) (McManus et al. 2016).
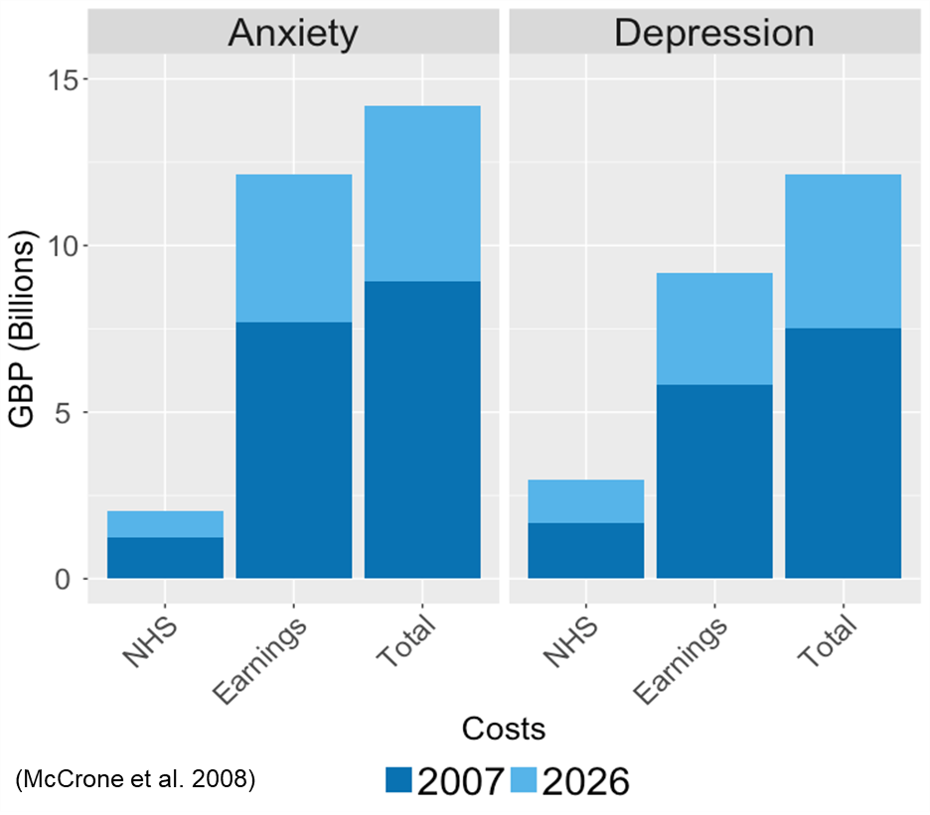
The high costs of anxiety and depression, on the health service, society and livelihood! Data from: (Mccrone et al. 2008). Currently, each disorder costs the UK £10billion pounds. It is estimated that savings could be around £25 million pounds per year if everyone received the treatment they needed.
There are various treatment options for anxiety and depressive disorders, which fall into two main classes. First, there are pharmacological treatments, including anxiolytics and anti-depressants. Second, and the focus of this blog, are psychological therapies, such as cognitive behavioural therapy (CBT), interpersonal therapy, and psychotherapy. In addition, well-being and self-help activities are becoming increasingly popular for individuals with milder symptoms. Depending on the severity and duration of illness, a doctor may recommend one or more of these approaches. However, not all of these treatments work for everyone. For example, cognitive behavioural therapy, an evidence-based goal-orientated treatment, only leads to a reduction of symptoms to below clinical threshold in around 50% of individuals. This means that for the other 50%, symptoms still cause impairment.
At present, many people face a “trial and error” approach where they experience several treatments that are not helpful before (hopefully) finding the right treatment(s) for them. A central goal of personalised medicine is to use patient information to help clinicians decide at the outset which treatment is most likely to work for whom. An important component of this is to identify which treatments are unlikely to work for any one individual, so they do not waste time, energy and personal resources on a treatment that probably won’t help. By making better predictions we hope to prevent additional patient distress, as well as time, money and resources.
Variation in response is observed in all varieties of treatment (each treatment works for some, but not for all). Indeed, much of the variance observed in therapeutic success may result from individuals not receiving the most appropriate type of treatment.
Most research investigating therapy response focuses on collecting and using clinical, social and demographic data to predict response. Numerous studies indicate that more severe symptoms, longer duration of illness, comorbidity, poor belief in treatment, poor adherence to treatment, poor learning ability and interpersonal difficulties lead to poorer treatment outcomes when treated with cognitive behavioural therapy. We expect that many more individual characteristics are likely to influence the success of treatment.
However, many such factors are likely to be genetically influenced. Twin studies show that complex traits such as anxiety, depression, learning ability and response to the environment, are around 30-50% heritable (Polderman et al. 2015). We therefore expect that response to therapy, like many environmental responses, will have a genetic component.
What do we know so far about genetic influences on psychological treatment response?
To date, the only evidence for a contribution of genetics to therapy response comes from candidate gene studies (Lester and Eley 2013). However, many candidate gene associations have failed to replicate. Therefore, the genetic effects that influence response to therapy are likely to be individually small and dispersed across the genome. This is the case with most other complex traits, where no single genetic variant has a substantial impact. This means that analyses should focus on genome-wide variation.
The first genome-wide therapygenetics investigation examined response to CBT in ~1,000 children with anxiety disorders (Coleman et al. 2016). No genome-wide significant genetic associations with treatment response were detected. However, as progress in genome-wide studies of schizophrenia and depression has shown, tens (if not hundreds) of thousands of individuals are required to detect single genetic variants associated with a complex trait. Nonetheless, these results provide more evidence that response to therapy is not influenced by common genetic variants with large effects.
However, other approaches, aggregating genetic variation at the level of genes, pathways and genome-wide polygenic scores can help to improve statistical power to detect variation associated with therapy response.
A second study (in the same child sample) examined the effects of a polygenic score for environmental sensitivity. This score was then used to test whether genetic propensity for environmental sensitivity predicted response to CBT (Keers et al. 2016). The study found that the environmental sensitivity polygenic score was positively associated with response to individual CBT (high intensity), but negatively associated with the brief parent led CBT (lower intensity).
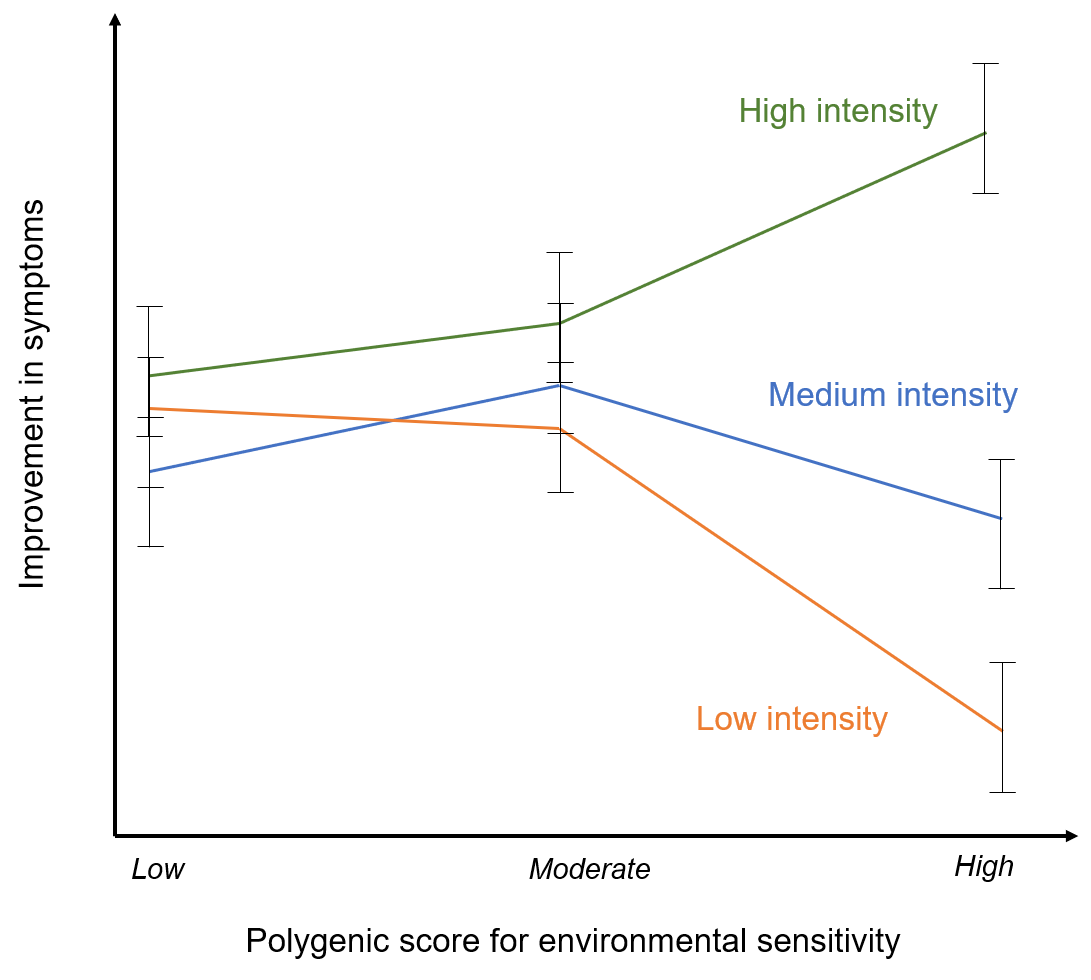
Effects of the low, moderate and high polygenic environmental sensitivity scores on change in anxiety symptoms for individuals treated with individual CBT, group CBT and brief parent-led CBT (adapted from: Keers et al. 2016). Here individuals who are more sensitive to the environment respond much more favourably to high intensity treatments than to low intensity treatments. In contrast, those with lower environmental sensitivity polygenic scores show similar rates of response for both the high and low intensity treatment formats (Keers et al. 2016).
Those individuals with a high polygenic score for responsivity to the environment responded most positively to the most intense type of treatment. This might suggest that individuals that are more genetically sensitive to the environment may be more likely to develop anxiety disorders under environmental stress but, may also benefit substantially from more intense treatment strategies.
While this finding is in need of replication, it clearly demonstrates how polygenic scores might be used to identify which individuals will benefit from which treatment.
Genome-wide polygenic scores add up the small effects of thousands of common, genetic variants into a single score for an individual for a specific trait. These scores borrow power from large population-based discovery studies, which can be applied to investigate the genetic overlap between the discovery trait and a trait of interest, i.e. treatment response. Larger genome-wide studies of behaviour and psychopathology will provide more accurate polygenic scores. High powered polygenic scores are already beginning to explain >10% of the variance in related traits (Selzam et al. 2016; Vassos et al. 2017; Lee et al. 2018).
Where we’re headed..
Several exciting papers in this emerging field are currently on their way through the publication process. We will update this blog when they come out, so watch this space!
The value of therapygenetics is dependent upon identifying genetic variance that can be useful to doctors – to help them choose the most favourable treatment for each patient. This work will require much larger samples than presently available to detect genetic effects. So if you’re a researcher and you have relevant data, please do get in touch! Or if you’re a patient who has experienced clinical levels of anxiety and depression and would like to take part in research (whether you have received treatment or not) you could join our Genetic Links to Anxiety and Depression (GLAD) Study.
References

